Computational
Design Platform
GPU-accelerated protein design that generates, scores, and validates thousands of candidates before anything touches your bench.
End-to-End Design Pipeline
Real Data, Not Mockups
These are actual outputs from the platform — not marketing graphics. Every project delivers ranked candidates with binding predictions, structural models, and developability scores.
The platform dashboard shows binding metrics across targets, score distributions, developability analysis, and design tool utilization. What you see here is what you get as a deliverable.
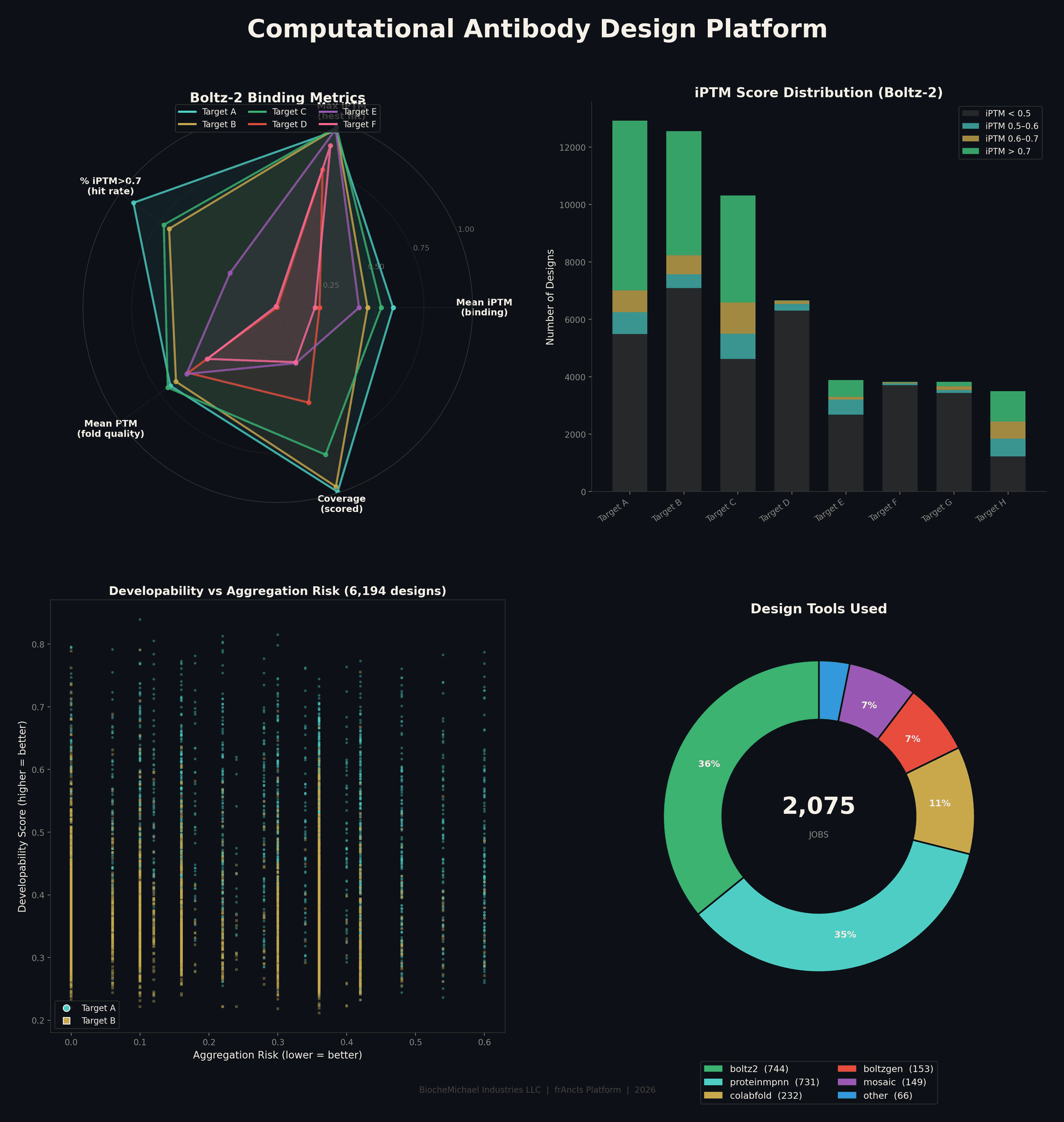
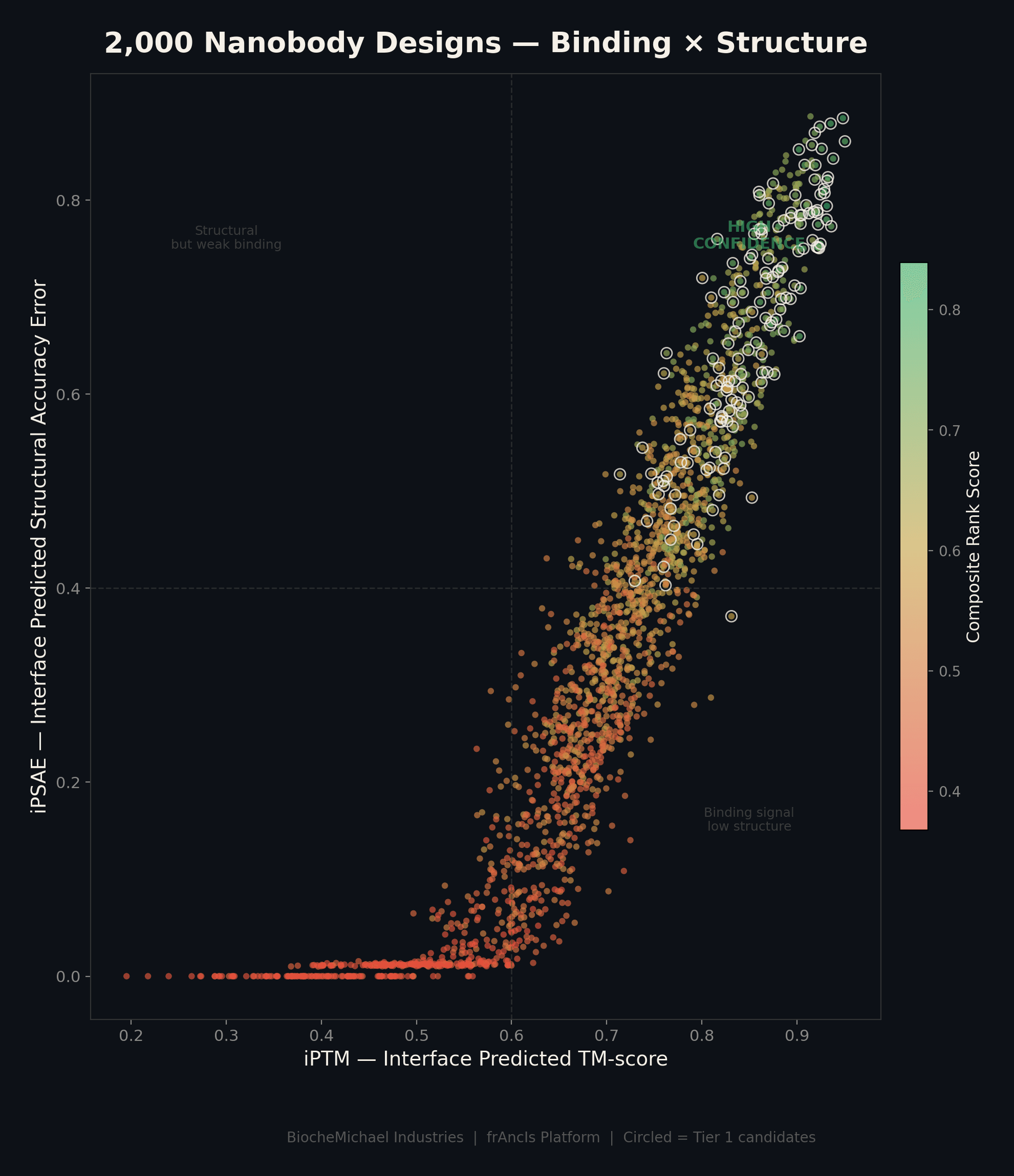
Screen 2,000 Candidates at Once
Every candidate is scored on two axes: predicted binding strength (iPTM) and structural accuracy (iPSAE). The top-right corner is where the best designs live — strong binding and confident structure. Only the top performers make the shortlist.
This is one target, one run. The platform generates and scores this volume routinely, giving you a diverse pool of candidates to choose from — not just one or two shots in the dark.
Custom-Trained AI Models
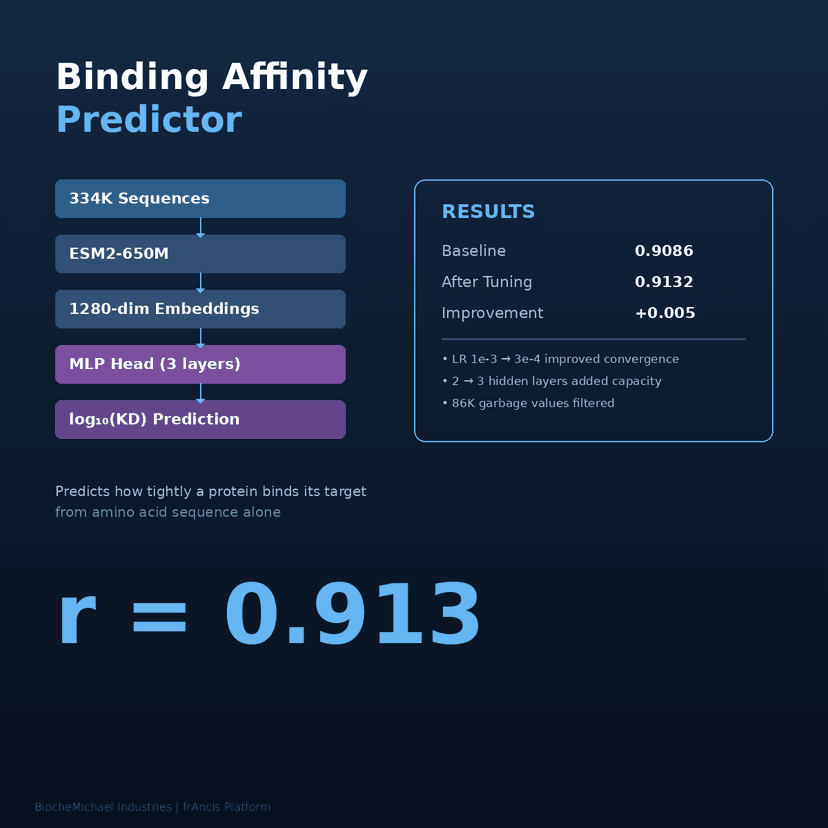
The binding affinity predictor is trained on 134K experimental sequences and achieves r = 0.913 correlation with measured KD values. It predicts how tightly a protein binds its target from amino acid sequence alone — no structure needed.
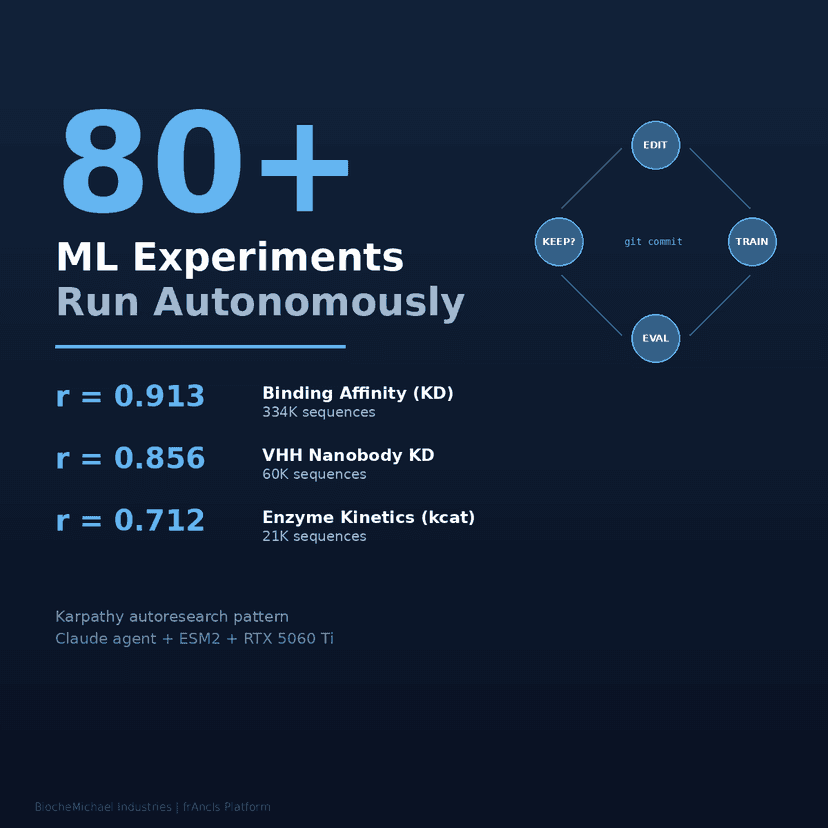
The platform continuously improves. An automated research system runs 80+ ML experiments autonomously, testing architectures, hyperparameters, and training strategies to push prediction accuracy higher.


Platform Capabilities
Every stage of the design pipeline, from target to candidate.
De Novo Backbone Generation
Diffusion models generate novel protein backbones conditioned on your target's binding surface. Not homology modeling — genuinely new structures designed from scratch.
Inverse Folding
Given a backbone, sequence design algorithms find amino acid sequences that fold into the desired structure. Multiple sequences per backbone, optimized for expression and stability.
AI Binding Prediction
Custom ML models trained on experimental binding data predict binding affinity for every candidate. Validated against lab measurements with strong correlation.
Structure Validation
Every candidate is folded in silico and checked for structural confidence. Designs that don't fold well computationally don't make the shortlist.
Automated Screening
The full pipeline runs autonomously — generate, score, validate, rank. Thousands of candidates processed in hours, not weeks.
Gradient Optimization
For lead optimization, gradient-based methods backpropagate through structure prediction to directly optimize binder sequences for affinity and stability.
Epitope Diversity Built In
The platform doesn't just find one binding mode — it explores the target surface and generates candidates across multiple distinct epitopes. This gives you backup strategies and reduces the risk of a single point of failure.
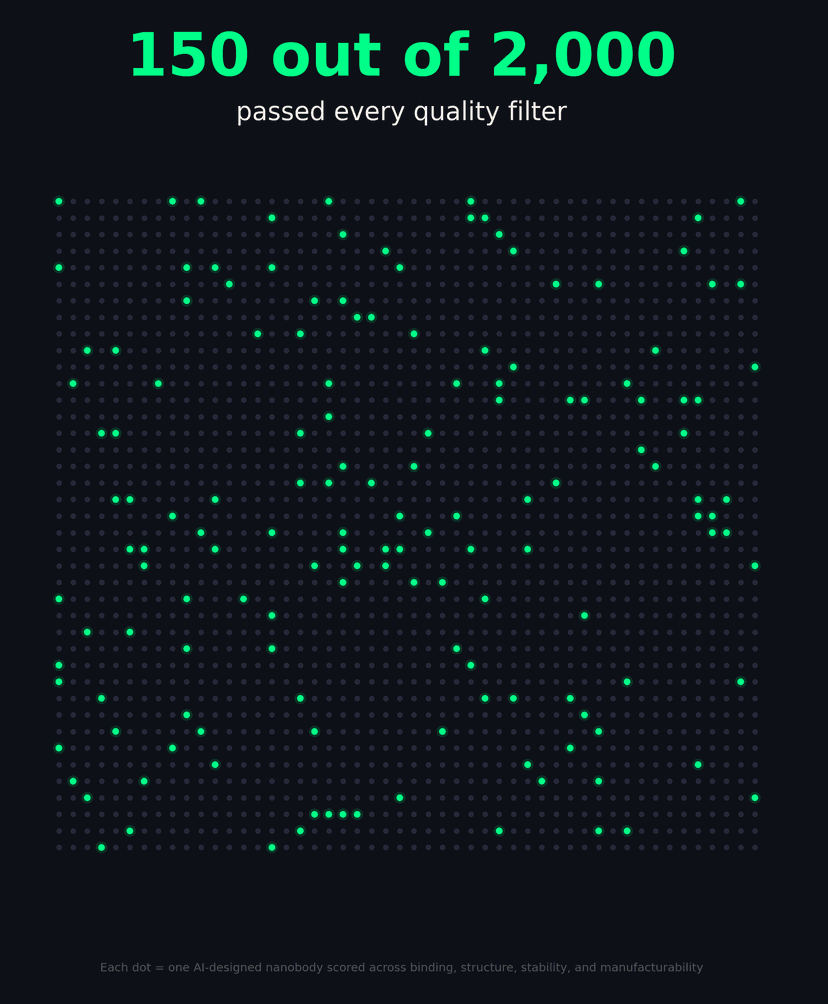
In one campaign: 270 designs across 5 distinct binding sites, with 150 out of 2,000 passing every quality filter — binding, structure, stability, and manufacturability.
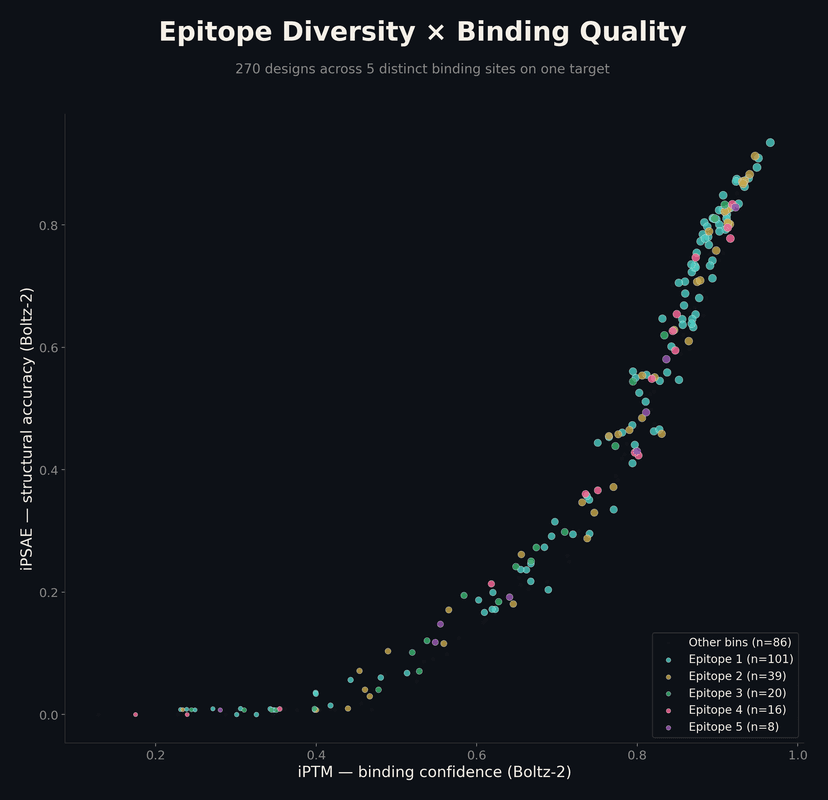

Want to See What This Can Do for Your Target?
Book a free 30-minute call. Bring your target structure or sequence—I'll give you an honest assessment.