Computational
Protein Design
Leveraging computational tools to inform and accelerate protein engineering. Structural modeling, binding prediction, and in silico screening.
Screen Thousands Before You Touch a Pipette
Traditional discovery screens hundreds of clones in the lab to find a handful of hits. Computational design flips that ratio—generate thousands of candidates in silico, rank them by predicted binding affinity and structural quality, and only synthesize the top performers.
I use GPU-accelerated structure prediction (AlphaFold2, Boltz-2, ESMFold), de novo backbone generation (RFDiffusion, BoltzGen), and custom ML models trained on experimental binding data to design and score protein binders. Every candidate gets a predicted binding affinity, fold confidence score, and developability assessment before it reaches your shortlist.
This isn't a black box. You get ranked candidate lists with full scoring data, structural models, and a clear rationale for every design decision. I work with your team to interpret results and prioritize what goes into the lab.
What I Deliver
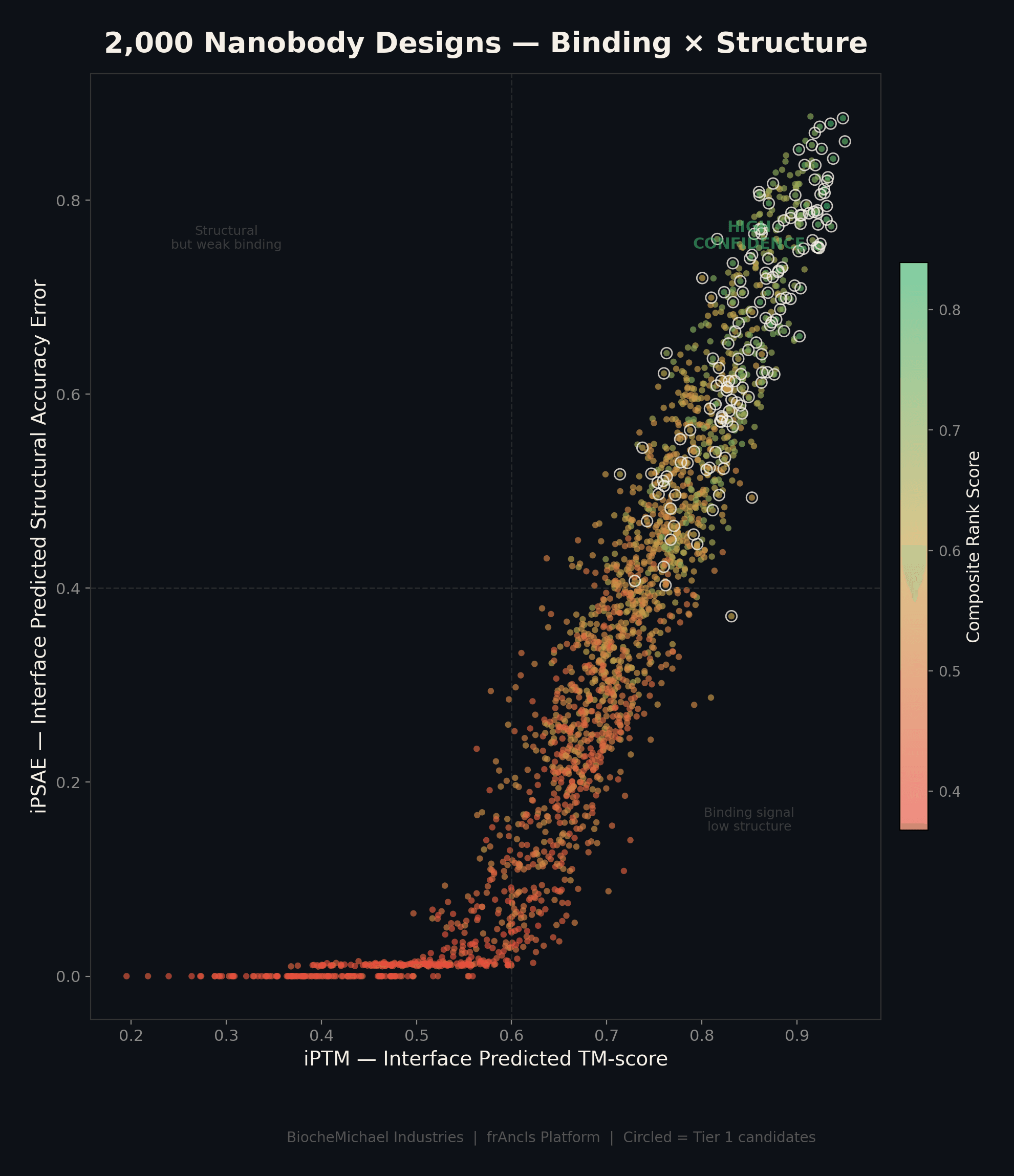
Computational Design Pipeline
Want to See What Computational Design Can Do for Your Program?
Book a free 30-minute call. Bring your target—I'll tell you what's possible.